Charles Margossian, Aki Vehtari, Daniel Simpson, Raj Agrawal write:
Gaussian latent variable models are a key class of Bayesian hierarchical models with applications in many fields. Performing Bayesian inference on such models can be challenging as Markov chain Monte Carlo algorithms struggle with the geometry of the resulting posterior distribution and can be prohibitively slow. An alternative is to use a Laplace approximation to marginalize out the latent Gaussian variables and then integrate out the remaining hyperparameters using dynamic Hamiltonian Monte Carlo, a gradient-based Markov chain Monte Carlo sampler. To implement this scheme efficiently, we derive a novel adjoint method that propagates the minimal information needed to construct the gradient of the approximate marginal likelihood. This strategy yields a scalable method that is orders of magnitude faster than state of the art techniques when the hyperparameters are high dimensional. We prototype the method in the probabilistic programming framework Stan and test the utility of the embedded Laplace approximation on several models, including one where the dimension of the hyperparameter is ∼6,000. Depending on the cases, the benefits are either a dramatic speed-up, or an alleviation of the geometric pathologies that frustrate Hamiltonian Monte Carlo.
They conclude:
Our next step is to further develop the prototype for Stan. We are also aiming to incorporate features that allow for a high performance implementation, as seen in the packages INLA, TMB, and GPstuff. Examples includes support for sparse matrices required to fit latent Markov random fields, parallelization and GPU support. We also want to improve the flexibility of the method by allowing users to specify their own likelihood. In this respect, the implementation in TMB is exemplary. It is in principle possible to apply automatic differentiation to do higher-order automatic differentiation and most libraries, including Stan, support this; but, along with feasibility, there is a question of efficiency and practicality. The added flexibility also burdens us with more robustly diagnosing errors induced by the approximation. There is extensive literature on log-concave likelihoods but less so for general likelihoods. Future work will investigate diagnostics such as importance sampling, leave-one-out cross-validation, and simulation based calibration.
One thing I can’t quite figure out from skimming the paper is whether the method helps for regular old multilevel linear and logistic regressions, no fancy Gaussian processes, just batches of varying intercepts and maybe varying slopes. I guess the method will work in such examples; it’s just not clear to me how much of a speed improvement you’d get. This is an important question to me because I see these sorts of problems all the time.
I’m also wondering if some of the computation could be improved by including stronger priors on the hyperparameters. Again, that’s an idea that’s been coming up a lot lately, in a wide range of applications.
Finally, I’m wondering how much parallelization is going on. Is this new algorithm faster because it requires fewer computations or because it is more parallelizable so you can get wall-time improvements by plugging in more processors? Either way is fine; I’d just like to have a better sense of how the method is working and where the speedup is coming from:

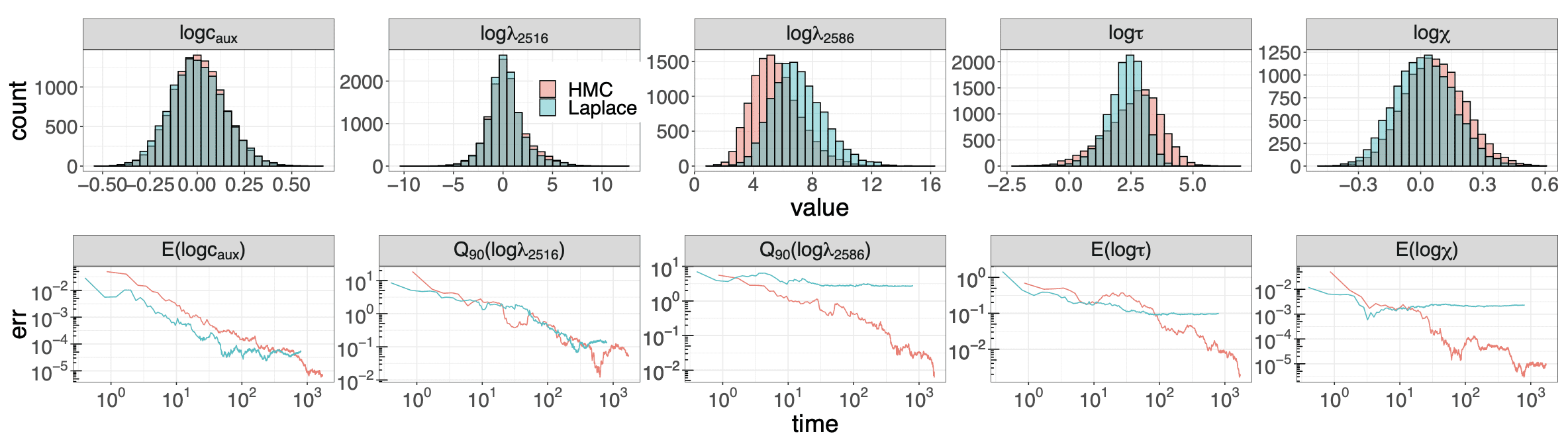
The y-axis of this graph should be on the log scale, but whatever.
lol @ the y-axis should be on the log scale snipe.. come on Andrew.. let them have this.. it seems great
https://www.medrxiv.org/content/10.1101/2020.05.13.20101253v2.full.pdf
Hi Andrew. Have you read this article but Ioannidis? I believe it came out 2 days ago. Would love your feedback on it. I´ve been following both you and Ioannidis during this crisis.
Best,
Ben
Ben:
Someone will have to look at all the individual studies that go into that report.
So Ioannidis cites the Imperial College estimate of 2 million deaths in US in absence of intervention in a way that suggests he thinks it was way out of the ballpark. Certainly he thinks his work on IFR is inconsistent with that. But here’s the deal- we are now ~110K deaths. Let’s say we care about scenario where 80% of population gets infected. Based on seroprevalence here it’s hard to believe we are nationally much more than 5%. Could even be lower. Multiply 110K by 16. That ballpark remains reasonable IMO and is vastly higher than seasonal influenza contra his earlier assertions.
OT, but amusing to say the least, will Alexie and Doc Walker be joining the fight? :)
This article says artificial brains need sleep too!
Hi Andrew,
> One thing I can’t quite figure out from skimming the paper is whether the method helps for regular old multilevel linear and logistic regressions
Yes, you can apply the method to multilevel models. The second example in the paper is a multilevel model with a logistic regression (and a horseshoe prior). But you’re right: you don’t always get a speed improvement. Sometimes the method can be slower because it’s expensive to recast the model as a latent Gaussian model and deal with the marginal covariance matrix. You can expect a speed up if the dimension of the of the latent variable is much higher than that of the hyperparameter.
Speedup is however not the only benefit. The latent variable, through its interaction with the data and the hyperparameter, produces a geometry that’s difficult to deal with (e.g. funnel shapes and others). If you marginalize out the latent variables, you remove the problem. In examples 2 and 3, you don’t get an important computational speedup. But you get rid of divergent transitions. With Stan’s HMC, it is extremely difficult to do this; it took me 4 – 5 runs (each 6 – 10 hours long) to — almost! — get rid of divergences; after non-centering, adjusting the target step size and the term buffer. With the embedded Laplace approximation, I could use Stan’s default tuning parameters. There is however a trade-off: the approximation introduces an error. It is marginal for a Poisson likelihood (example 1), more important for a Bernoulli likelihood — whether it matters or not depends on the quantities of interest and your goals as a modeler.
> I’m also wondering if some of the computation could be improved by including stronger priors on the hyperparameters.
Yes and no. From a computational perspective, you want to make the curvature more even and it’s not always clear that a stronger prior achieves this. But I agree that some computational issues can be resolved by including more information in the problem, notably in the form of better constructed priors.
> Finally, I’m wondering how much parallelization is going on
None. Note that the paper compares different things and it’s important to distinguish the different computer experiments. Plot 1 (at the bottom of the post, figure 1 in the paper) examines the differentiation time of the marginal likelihood using two methods. One is the algorithm in Rasmussen & Williams (2006), implemented in the package GPStuff. The other one is home brewed. The benchmark algorithm is really clever; but it was designed for problems where the derivative of the covariance matrix, K, was readily available and the dimension of the hyperparameter low. We had problems where neither was true. You can get rid of the first constraint by using automatic differentiation to differentiate K. It works but it doesn’t scale because computing the tensor of derivatives of K becomes very expensive, both with forward and reverse mode autodiff. So how do you bypass that? You don’t compute the derivatives of K.
That’s what I would call the wisdom of autodiff: don’t explicitly compute the Jacobian of intermediate operations. The appendix walks you through the idea. It’s the same principle we’re using to improve ODEs in Stan and that we used for HMMs. And there are many more areas in Stan — and beyond — where this idea can be applied. It’s nice to demonstrate how important this principle is and really hammer down the point that autodiff isn’t a simple application of the chain rule.
Sorry for the long comment: we can discuss more in person!
Then can we expect the speedup when
1. hyperparameter has high dimension (second constraint of GPStuff alg.)
2. latent variable dimension is much higher than that of the hyperparameter
In other words, when p is high but n is even higher than p (from your paper)?
To be clear, dim( φ ) is large and dim(θ) ≫ dim( φ )?
I got confused as n is used both as a dimension of latent variable(3.1) and the number of observation.
One more question:
The current dataset has dim(θ) = 102, dim( φ ) = 5966. Would the speedup be more salient with higher dim(θ)?
Hi Hyunji, that’s a good question.
As you probably noted, we didn’t test a case where p is large and n is even larger. So it’s certainly worth trying it out, if you have a model with this configuration.
My intuition is that yes, you will get a speedup. I’m quite confident of that if p = 20 or p = 100, and n = 1,000. But it’s worth studying how the geometry of the posterior changes in these “doubly” high-dimensional cases.