I’m implementing the matrix normal distribution for Stan, which provides a multivariate density for a matrix with covariance factored into row and column covariances.
The motivation
A new colleague of mine at Flatiron’s Center for Comp Bio, Jamie Morton, is using the matrix normal to model the ocean biome. A few years ago, folks in Richard Bonneau‘s group at Flatiron managed to extend the state of the art from a handful of dimensions to a few dozen dimensions using Stan, as described here
- Tarmo Äijö, Richard Bonneau, and Harri Lähdesmäki. 2018. Generative Models for Quantification of DNA Modifications. Data Mining for Systems Biology.
That’s a couple orders of magnitude short of the models Jamie would like to fit. So instead of Stan, which he’s used before and is using for other smaller-scale problems, he’s turning to the approach in the following paper:
- Justin D. Silverman, Kimberly Roche, Zachary C. Holmes, Lawrence A. David, and Sayan Mukherjee. Bayesian Multinomial Logistic Normal Models through Marginally Latent Matrix-T Processes. arXiv.
I’d just like to add matrix normal to Stan and see if we can scale up Äijö et al.’s results a bit.
The matrix normal
The density is defined for an
- $latex N \times P$ observation matrix $latex Y$
with parameters
- $latex N \times P$ mean matrix $latex M$,
- positive-definite $latex P \times P$ column covariance matrix $latex U$, and
- positive-definite $latex N \times N$ row covariance matrix $latex V$
as
$latex \displaystyle \textrm{matrix\_normal}(Y \mid M, U, V) = \frac{\displaystyle \exp\left(-\frac{1}{2} \cdot \textrm{tr}\left(V^{-1} \cdot (Y – M)^{\top} \cdot U^{-1} \cdot (Y – M)\right)\right)}{\displaystyle (2 \cdot \pi)^{{N \cdot P} / {2}} \cdot \textrm{det}(V)^{{N} / {2}} \cdot \textrm{det}(U)^{{P} / {2}}}.$
It relates to the multivariate normal through vectorization (stacking the columns of a matrix) and Kronecker products as
$latex \textrm{matrix\_normal}(Y \mid M, U, V) = \textrm{multi\_normal}(\textrm{vec}(Y) \mid \textrm{vec}(M), V \otimes U).$
After struggling with tensor differentials for the derivatives of the matrix normal with respect to its covariance matrix parameters long enough to get a handle on how to do it but not long enough to get good at it or trust my results, I thought I’d try Wolfram Alpha. I could not make that work.
matrixcalculus.org to the rescue
After a bit more struggling, I entered the query [matrix derivative software] into Google and the first hit was a winner:
- Matrix and vector derivative caclulator at matrixcalculus.org
This beautiful piece of online software has a 1990s interface and 2020s functionality. I helped out by doing the conversion to log scale and dropping constant terms,
$latex \begin{array}{l}\displaystyle\log \textrm{matrix\_normal}(Y \mid M, U, V) \\[8pt] \qquad = -\frac{1}{2} \cdot \left(\textrm{tr}\!\left[V^{-1} \cdot (Y – M)^{\top} \cdot U^{-1} \cdot (Y – M)\right] – N \cdot \log \textrm{det}(V) – P \cdot \log \textrm{det}(U) \right) + \textrm{const}.\end{array}$
Some of these terms have surprisingly simple derivatives, like $latex \frac{\partial}{\partial U} \log \textrm{det}(U) = U^{-1}$. Nevertheless, when you push the differential through, you wind up with tensor derivatives inside that trace which are difficult to manipulate without guesswork at my level of matrix algebra skill.
Unlike me, the matrix calculus site chewed this up and spit it out in neatly formatted LaTeX in a split second:
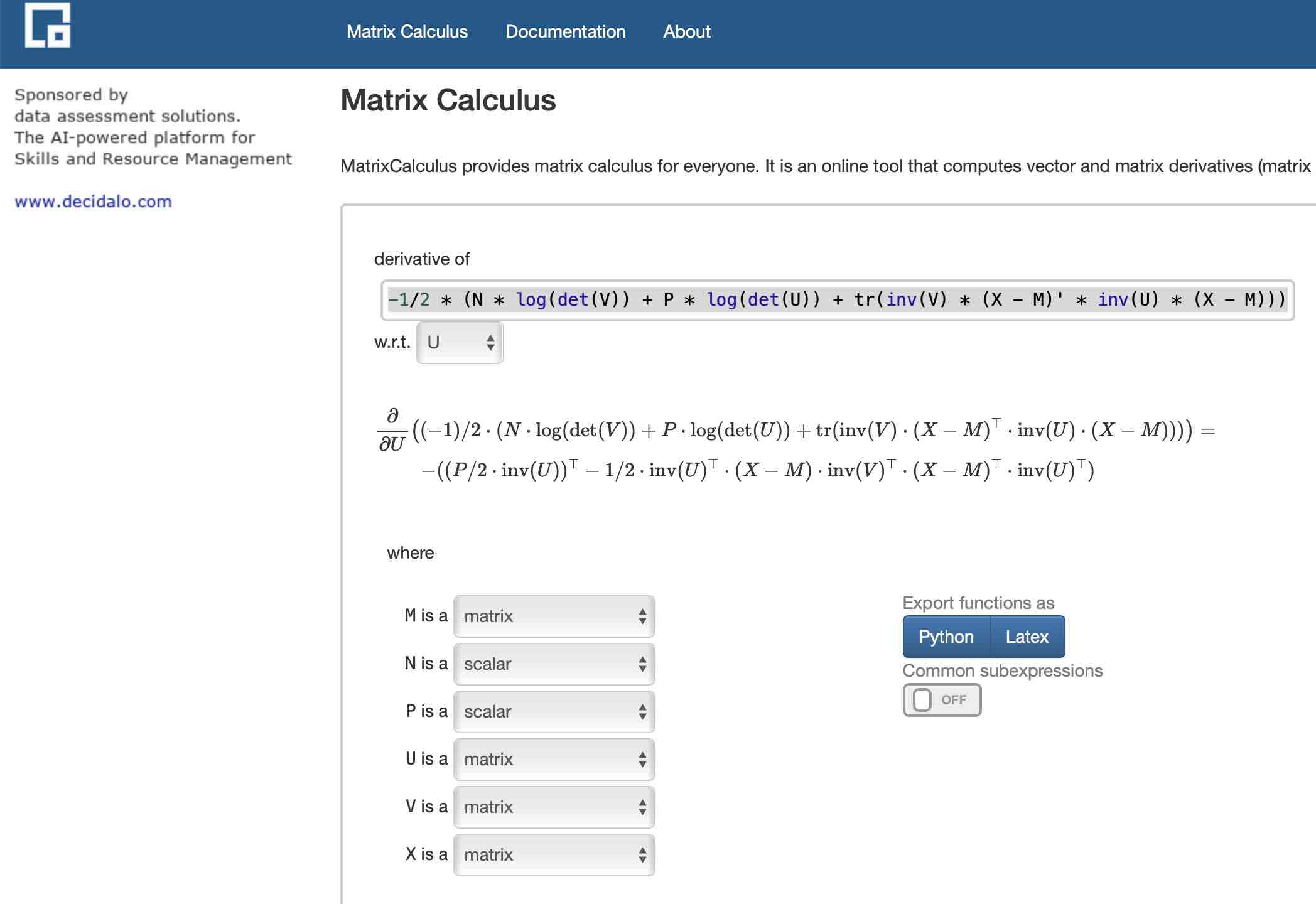
Despite the simplicity, this is a really beautiful interface. You enter a formula and it parses out the variables and asks you for their shape (scalar, vector, row vector, or matrix). Another win for delcaring data types in that it lets the interface resolve all the symbols. There may be a way to do that in Wolfram Alpha, but it’s not baked into their interface.
The paper
There’s a very nice paper behind that explains what they did and how it relates to autodiff.
- S. Laue, M. Mitterreiter, and J. Giesen. 2018. Computing Higher Order Derivatives of Matrix and Tensor Expressions. NeurIPS.
I haven’t digested it all, but as you may suspect, they implement a tensor algebra for derivatives. Here’s the meaty part of the abstract.
The framework can be used for symbolic as well as for forward and reverse mode automatic differentiation. Experiments show a speedup of up to two orders of magnitude over state-of-the-art frameworks when evaluating higher order derivatives on CPUs and a speedup of about three orders of magnitude on GPUs.
As far as I can tell, the tool is only first order. But that’s all we need for specialized forward and reverse mode implementations in Stan. The higher-order derivatives start from reverse node then nest in one more forward mode instances. I’m wondering if this will give us a better way to specialize fvar<var> in Stan (the type used for second derivatives).
Some suggestions for the tool
It wouldn’t be Andrew’s blog without suggestions about improving interfaces. My suggestions are to
- check the “Common subexpressions” checkbox by default (the alternative is nice for converting to code),
- stick to a simple checkbox interface indicating with a check that common subexpression sharing is on (as is, the text says “On” with an empty checkbox when it’s off and “Off” with an empty checkbox when it is on),
- get rid of extra vertical whitespace output at hte bottom of the return box, and
- provide a way to make the text entry box bigger and multiple lines (as is, I composed in emacs and cut-and-paste into the interface), and
- allow standard multi-character identifiers (it seems to only allow single character variable names).
I didn’t try the Python output, but that’s a great idea if it produces code to compute both the function and partials efficiently.
Translating to the Stan math library
With the gradient in hand, it’s straightforward to define efficient forward-mode and reverse-mode autodiff in Stan using our general operands-and-partials builder structure. But now that I look at our code, I see our basic multivariate-normal isn’t even using that efficient code. So I’ll have to fix that, too, which should be a win for everyone.
What I’m actually going to do is define the matrix normal in terms of Cholesky factors. That has the huge advantage of not needing to put a whole covariance matrix together only to factor it (we need a log determinant). Sticking to Cholesky factors the whole way is much more arithmetically stable and requires only quadratic time to factor rather than cubic.
In terms of using the matrix derivative site, just replace U and V with (L_U * L_U’) and (L_V & L_V’) in the formulas and it should be good to go. Actually, it won’t because the parser apparently requires single letter variabes. So I wound up just using (U * U’), which does work.
Here’s the formula I used for the log density up to a constant:
-1/2 * (N * log(det(V * V')) + P * log(det(U * U')) + tr([inv(V * V') * (Y - M)'] * [inv(U * U') * (Y - M)]))
Because there’s no way to tell it that U and V are Cholesky factors, and hence that U * U’ is symmetric and positive definite, I had to do some reduction by hand, such as
inv(U * U') * U = inv(U')
That yields a whole lot of common subexpressions between the function and its gradient, and I think I can go a bit further by noting (U * U’) = (U * U’)’.
matrix_normal_lpdf(Y | M, U, V)
= -1/2 * (N * log(det(V * V'))
+ P * log(det(U * U'))
+ tr([inv(V * V') * (Y - M)'] * [inv(U * U') * (Y - M)]))
+ const
d/d.Y: -[inv(U * U') * (Y - M)] * inv(V * V')
d/d.M: -[inv(U * U') * (Y - M)] * inv(V * V')
d/d.U: -P * inv(U')
- inv(U * U') * (Y - M)] * [inv(V * V') * (Y - M)'] * inv(U')
d/d.V: -N * inv(V')
-[inv(V * V') * (Y - M)'] * [inv(U * U') * (Y - M)] * inv(V')
This is totally going in the book
Next up, I’d like to add all the multivariate densities to the following work in progress.
-
Bob Carpenter, Adam Haber, and Charles Margossian. 2020 (draft). The Automatic Differentiation Handbook. [GitHub link]
There’s a draft up on GitHub with all the introductory material and a reference C++ implementation and lots of matrix derivatives and even algebraic solvers and HMMs. Lots still to do, though.
We haven’t done the main densities yet, so we can start with multivariate normal and T with all four parameterizations (covariance, precision and Cholesky factors of same), along with the (inverse) Wisharts and LKJ we need for priors. This matrix calculus site’s going to make it easy to deal with all the Cholesky-based parameterizations.
If you’d like to help implementing these in Stan or in joining the effort for the handbook, let me know. I recently moved from Columbia University to the Flatiron Institute and I’m moving to a new email address: [email protected]. I didn’t change GitHub handles. Flatiron is great, by the way, and I’m still working with the Stan dev team including Andrew.
Hi Bob,
– U*U’ is always symmetric and positive semi-definite (if U is nonsingular, then it’s positive definite).
– It’s still cubic in N and P with cholesky factors. That’s because terms like inv(V * V’) * (Y – M)’ involve P solves (at O(P^2) each). Similar for the (U*U’) term (except replace N with P). So that matrix-matrix solve is approximately as expensive as the original Cholesky in terms of the operation count.
Like for our normal distribution, I’m assuming that U and V are full rank—the diagonals will be constrained to be strictly positive. But of course, that can fail numerically and we can wind up with zeros where we shouldn’t.
How does inv(V * V’) * (Y – M)’ involve P solves? V is just a matrix, so inv(V * V’) = inv(V’) * inv(V) involves just one inv(V) because inv(V’) = inv(V)’.
The matrix-matrix multiply will be cubic, but I don’t see why we need more than two solves, one for U and one for V.
The golden rule of numerical linear algebra is that you should only ever compute the inverse if you need it’s elements directly.
To get inv(V) the pseudocode goes
In: Matrix V
Out: Matrix Vinv
for p in 1:P
solve V * Vinv[:, p] = Id[:,p];
where Id is the identity matrix.
This means that you do P solves already to compute Vinv *and then* do a cubic matrix-matrix multiply to compute inv(V) * B (here B is some matrix of the right shape. I think it’s (Y-M)’ in your code). Instead, you could do this
In: Matrix V, B
Out: Matrix Vinv_times_B
for p in 1:P
solve V * Vinv_times_B[:, p] = B[:,p];
which avoids the extra storage and the extra matrix-matrix product. (In your example, there would be two matrix-matrix products, so even more saving)
Everything’s a Cholesky factor here in the real application. Will the inverse of a Cholesky factor be so unstable that we should do many more solves instead?
For the second example, why not compute inv(V) * B as V \ B with a single solve? Are the types of V and B not matrices?
What would be gained by using the explicit inverse of the cholesky factors? Parallelism?
We need the P * inv(V * V’) and N * inv(U * U’) terms for derivatives w.r.t. V and U. The only way I could see to fully get away from calculating those inverses is to postpone them until the adjoint-Jacobian product gets calculated and there’s an adjoint matrix to multiply.
I think we’re taking across each other. The first piece of pseudocode is how inv(V) is computed (through a lot of solves).
The second piece of pseudocode is how V \ B is evaluated.
Using inv(V) explicitly takes more operations and requires more memory than doing V\B.
Maybe a different way to put it is that inv(V) * B is roughly equivalent to (V \ Id) * B, where Id is the correct identity matrix.
OK. We only need the compound terms like inv(U * U’) * (Y – M). And see the above (below?) comment about inverses. We could get away from those with a custom chain() instead of using operands-and-partials. That’d also let us be more conservative with memory allocation because we don’t need to store the Jacobians in the forward pass and they’re just temporaries in the reverse (or even implicit if we unfold).
Also, what’s the motivation for reimplementing this over just using syntactic sugar for the vectorized one? Is it going to be more efficient? Because otherwise it seems like a lot of extra code in the math library for something we can do by adding a bit of code to the compiler.
We don’t have a matrix normal implementation now other than for a full precision matrix. And that just autodiffs through the matrix algebra. I’m going to build one using operands-and-partials that’ll be much more efficient. Although still cubic, so it’ll wind up being something like O(N * P^2 + N^2 * P)—I didn’t bother working it out.
You mean why not convert to multi_normal using vec()? We don’t have a multi-normal that can do anything other than expand the Kronecker product, which wouldn’t be workable.
I also don’t understand how we could do this in the compiler. We can’t generate efficient autodiff there. Or at least we haven’t tried to go that far yet.
It should work directly by plugging into muti_normal_cholesky_lpdf if eigen has support for vec (which it does just by storing it column major) and a kronecker product (which is unsupported in Eigen 3.3.3, but it’s there). (Remember the cholesky of the kronecker product is the kronecker product of the choleskys)
If you’re willing to wait for static matrices, you should be able to do it really smoothly, otherwise it will probably be a bit awkward (because it will work better with a MatrixXd than with a MatrixXv).
If you make a git issue it would probably be a better place to talk about this.
Ugh. This ate my comment.
But eigen has the equivalent of Vec (it’s the internal storage! Good old column major!) and there’s an unsupported kroneckerProduct module, so it should be easy to just use those in the existing multi_normal_lpdf and multi_normal_cholesky_lpdf functions. The latter works because chol( kron(A, B)) = kron(chol(A), chol(B)).
This would obviously be more efficient for matrices of doubles (otherwise it’s a painful loop through the matrix of vars) or static var matrices, but it should avoid the need to do all the bespoke derivatives.
At least, it would be worth trying it out on the one you’ve got implemented to see if there’s a real performance gain to doing it all by hand.
(If you want to talk more about this, we should probably do it in the git repo, because I miss markdown!)
I don’t have anything implemented yet. I don’t see us getting all the Kronecker stuff in place for the multivariate normal any time soon. It’s a lot of expression templates and new functionality around Kronecker products. I think I can write a reverse-mode implementation in a couple of days, at least half of which will be testing we’ll need no matter how we implement it.
If I’m willing to go with storing the Jacobians (four matrices in, scalar out, so it’s only the size of the input), then dealing with containers of autodiff variables isn’t so bad. I can use operands-and-partials, which will handle both forward and reverse mode.
Once we get immutable (not static) matrices, we’ll be trying to push their use everywhere possible, and should even be able to figure out that a lot of matrices are immutable even if they aren’t declared as such.
Dear Bob,
I am the author of the online tool matrixcalculus.org. I am very happy that it is of some use to you!
I am eager to improve the interface and I will take your five suggestions into consideration. And I will also silently whine a little bit about your comment “1990s interface and 2020s functionality”. Really that bad? I guess its time for a fresh new look then.
Sören:
Coming from Bob, “1990s interface and 2020s functionality” is the ultimate compliment. If he’d said, “1990s functionality and 2020s interface”—that would’ve been bad news! (I speak as someone whose webpage has a 1990s look and feel.)
;-)
Mine, too!
Thanks again for the awesome tool. I personally like the affordances offered by standard interface components like the dropdown lists for the data type. It’s not really a ’90s interface on the back end in that each of the interface elements when updated updates the whole site. That you couldn’t do with a ’90s web form. I don’t like fancy Javascript web interfaces that don’t look like web pages and use non-standard components and are hard to figure out. But then I don’t think Apple’s devices are as usable as the folks in Cupertino do. I literally couldn’t figure out how to turn off my first iPod and had to look it up on the web (the answer was to hold down the play button until it turned off).
Yeah, and it was really nice of Apple to insist on a one-button mouse so it’s impossible to click the wrong button…but then give us the control, option, and command keys to modify the mouse click (or a keypress). You can’t click the wrong mouse button, but you can sure as hell hold down the wrong modifier key — I know because I do it all the time.
That said, most technology comes with a lot of conventions that need to be mastered. There’s a YouTube channel that features children of various ages being given some bit of old technology and challenged to use it: make a call with a dial telephone, use a Walkman to play a song and then advance to the next song, etc. These can be pretty amusing, really driving home the extent to which things that were “natural” to my generation were actually things that had to be learned and often practiced. (Even once a teen figured out how to use a telephone dial — he had seen it done in old movies — he had trouble moving his hand in the nice circle required to make it work).
Interesting post. I don’t have much experience with fitting matrix normal distributions because I tend to prefer to impose some kind of structure on the model.
However, I have a lot of experience with fitting multivariate normal distributions with Stan. There have been a lot of difficulties, primarily because the data that I am interested in (time series economic and financial series) tend to be highly correlated. I think the LKJ prior was more of a hindrance. I recall the one thing that ended up working well was breaking apart the multivariate distribution into individual variables and regression problems (e.g., fit univariate normal for variable 1, then regress variable i against variables 1:(i-1)).
It would be interesting to see some information about enhancements to the multi_normal (like how much it is sped up, though that has never really been my big issue).
People may also be interested in http://www.geno-project.org/ which allows you to differentiate an objective function with respect to matrices and vectors using a simple language. That is less useful for Bob’s handbook but maybe more useful for stuffing everything into a function that evaluates a log-kernel (which I mentioned to Bob on Discourse back on January 14th, along with matrixcalculus.org).
I must have never followed up the reference to matrixcalculus.org or I would’ve been raving about how cool it was then.
GENO is part of the same project. That was a lot of Python output, but presumably one piece is the objective function. You can download Python code that’ll evaluate derivatives through their simpler site, too.
You were (accurately) raving about how cool it was then. https://discourse.mc-stan.org/t/help-write-an-autodiff-handbook/12640/4?u=bgoodri
I was saying that it sounded cool from the description—I don’t think I ever checked it out. If I did, I’ve completely forgotten about it or didn’t realize the ramifications at the time. I often don’t understand solutions to problems when they’re presented before I fully understand the problem.
Every time I go into Discourse these days, I get completely overwhelmed. Which then discourages me from visiting, which creates a vicious cycle where it takes even longer when I do get to it.
Bob,
This is very cool! Would this fast matrix-normal distribution make it easier to implement the array-normal distribution developed by Peter Hoff? (cite: https://projecteuclid.org/euclid.ba/1339612040)
Same idea, but Hoff’s has more dimensions of variation; from the abstract:
You might find these references handy for your book.
@incollection{giles2008collected,
title={Collected matrix derivative results for forward and reverse mode algorithmic differentiation},
author={Giles, Mike B},
booktitle={Advances in Automatic Differentiation},
pages={35–44},
year={2008},
publisher={Springer}
}
@article{giles2008extended,
title={An extended collection of matrix derivative results for forward and reverse mode automatic differentiation},
author={Giles, Mike},
year={2008},
publisher={Unspecified}
}